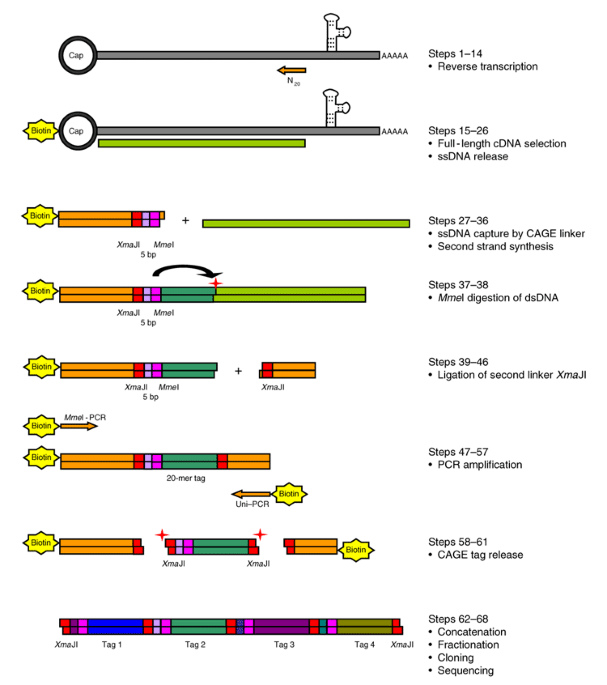
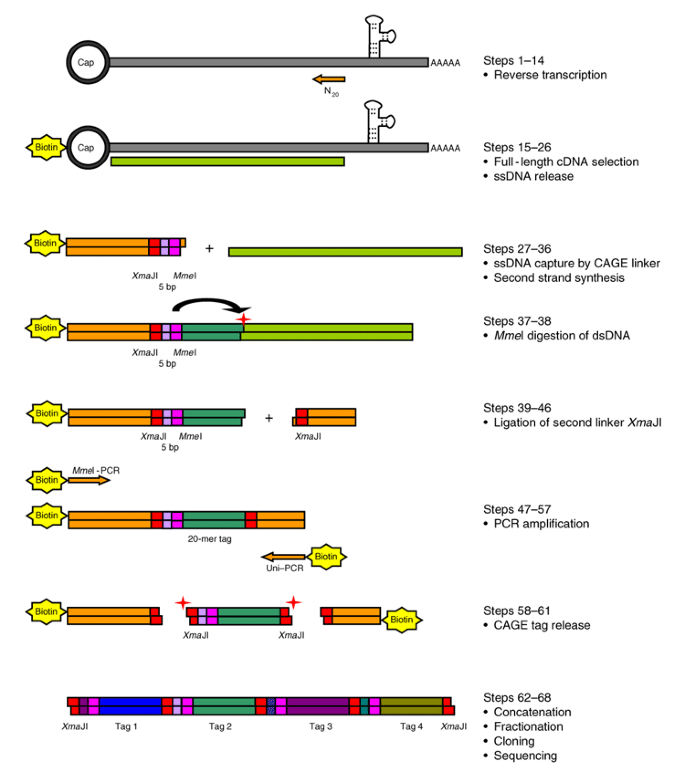
Solving the transcription start site identification problem with ADAPT-CAGE: a Machine Learning algorithm for the analysis of CAGE data | Scientific Reports

Integration of Single-Cell RNA- and CAGE-seq Reveals Tooth-Enriched Genes - Y. Chiba, K. Yoshizaki, T. Tian, K. Miyazaki, D. Martin, , Genomics and Computational Biology Core, Genomics and Computational Biology Core, K.
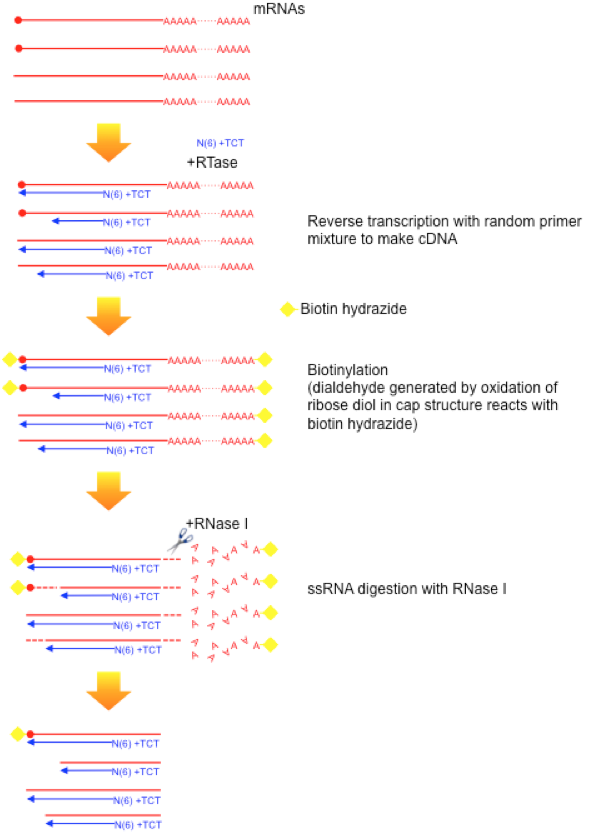
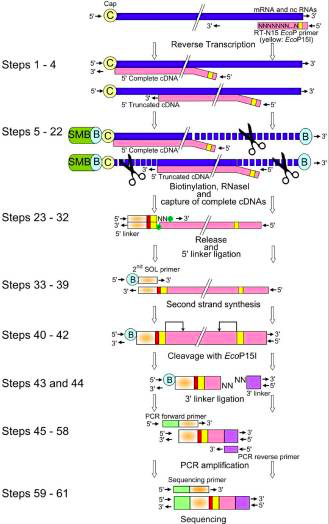
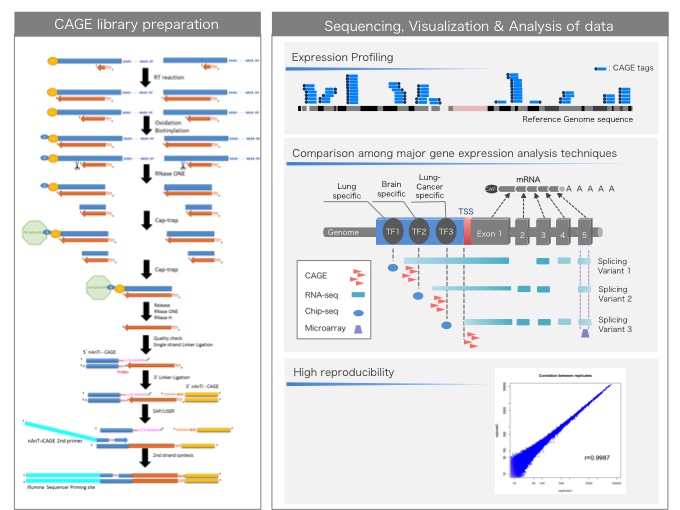
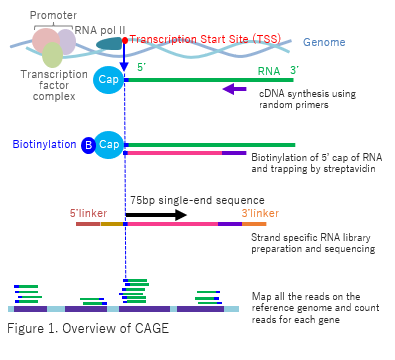
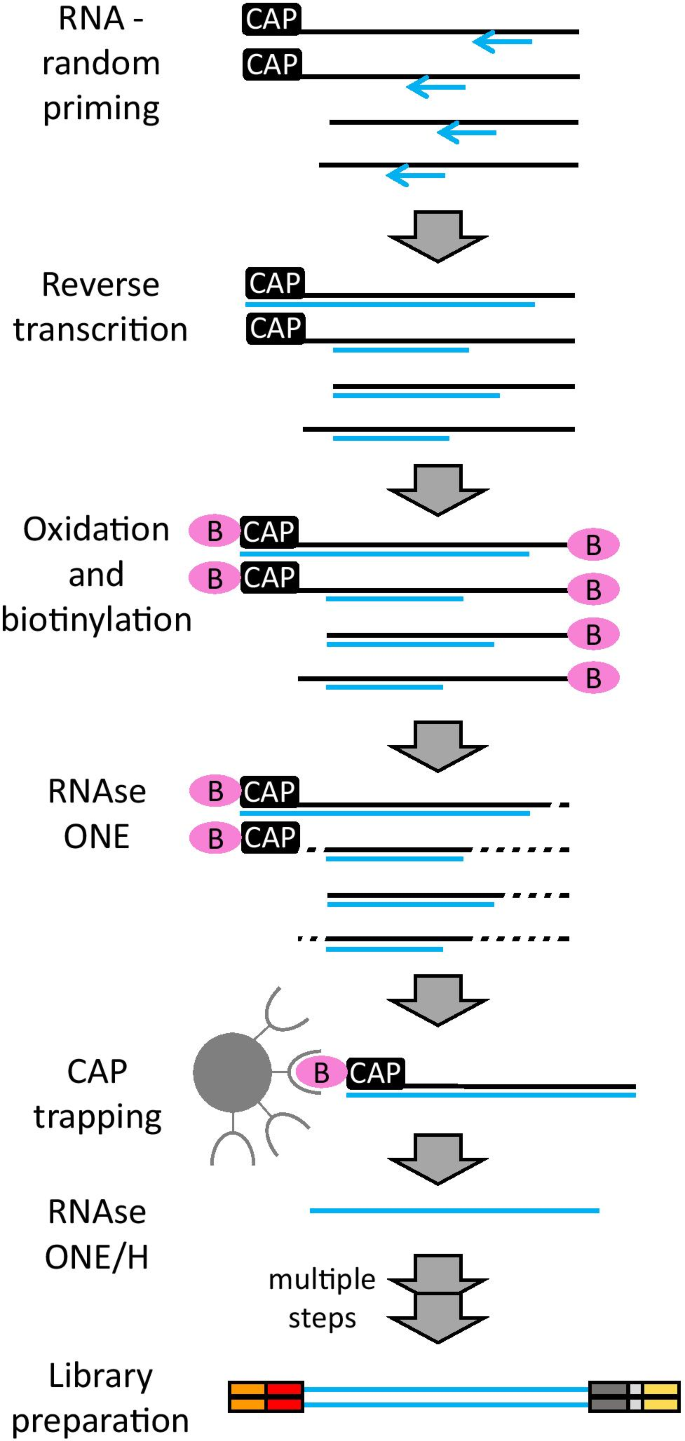
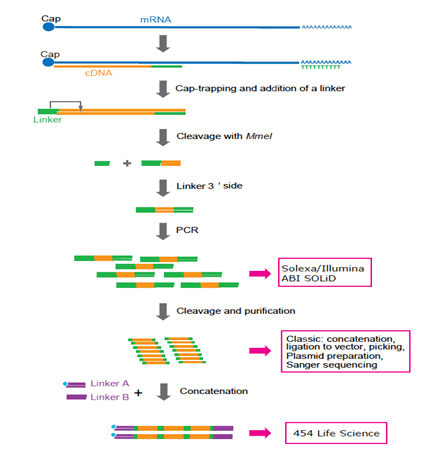
CAGE : A method for genome-wide identification of transcription start sites | 理化学研究所 ライフサイエンス技術基盤研究センター
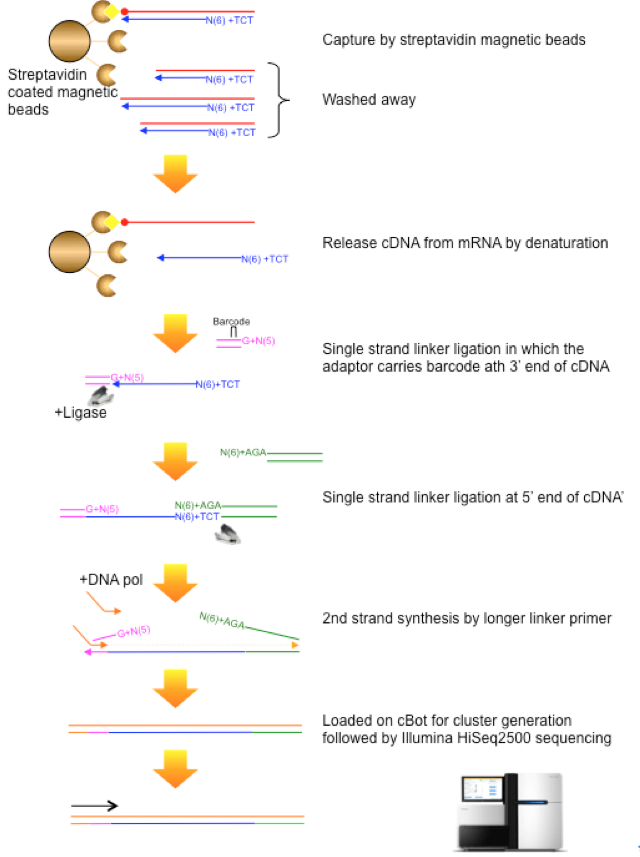
CAGE : A method for genome-wide identification of transcription start sites | 理化学研究所 ライフサイエンス技術基盤研究センター
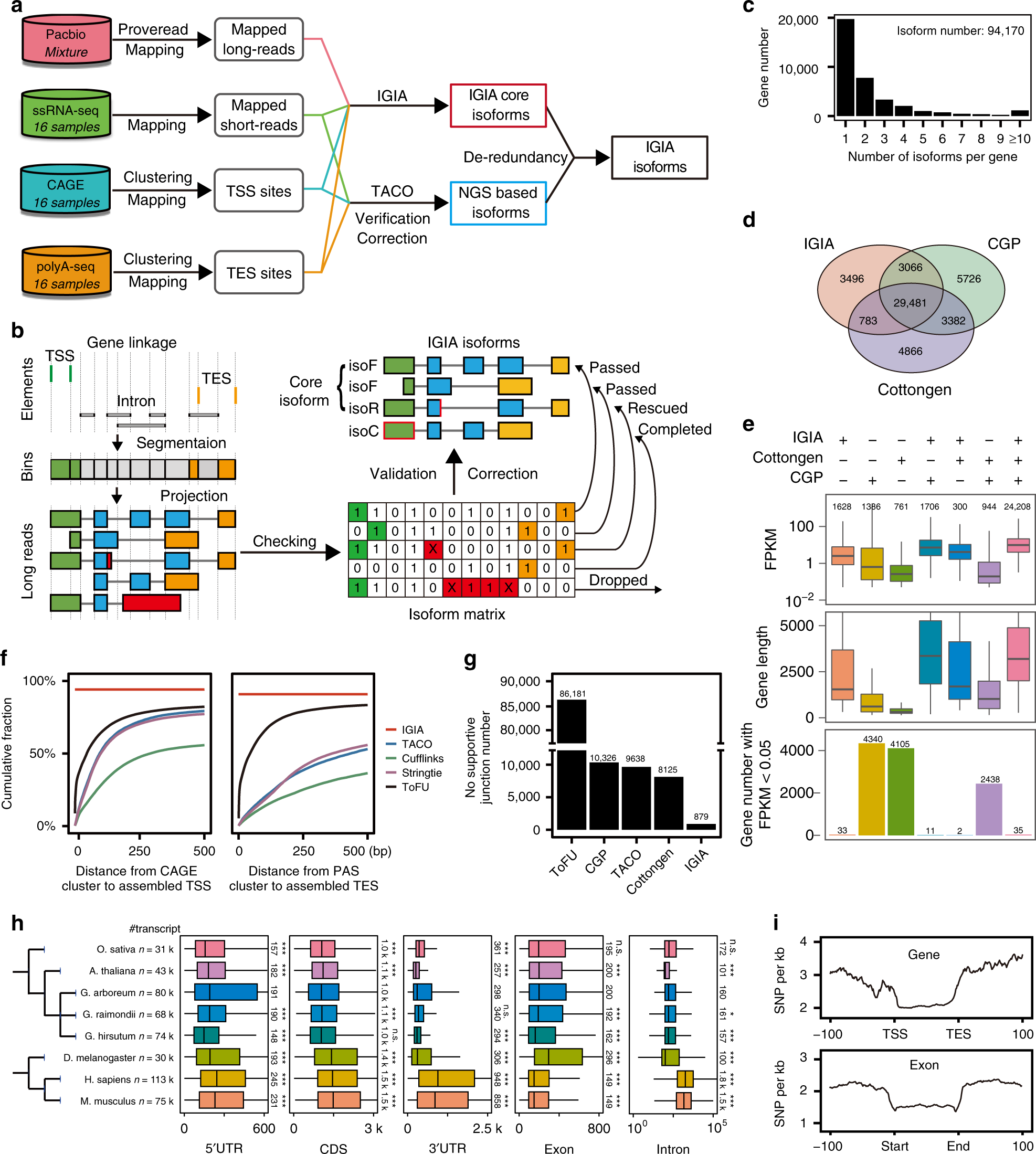
Multi-strategic RNA-seq analysis reveals a high-resolution transcriptional landscape in cotton | Nature Communications

Development of NET-CAGE a, Schematic of nascent RNA isolation in the... | Download Scientific Diagram

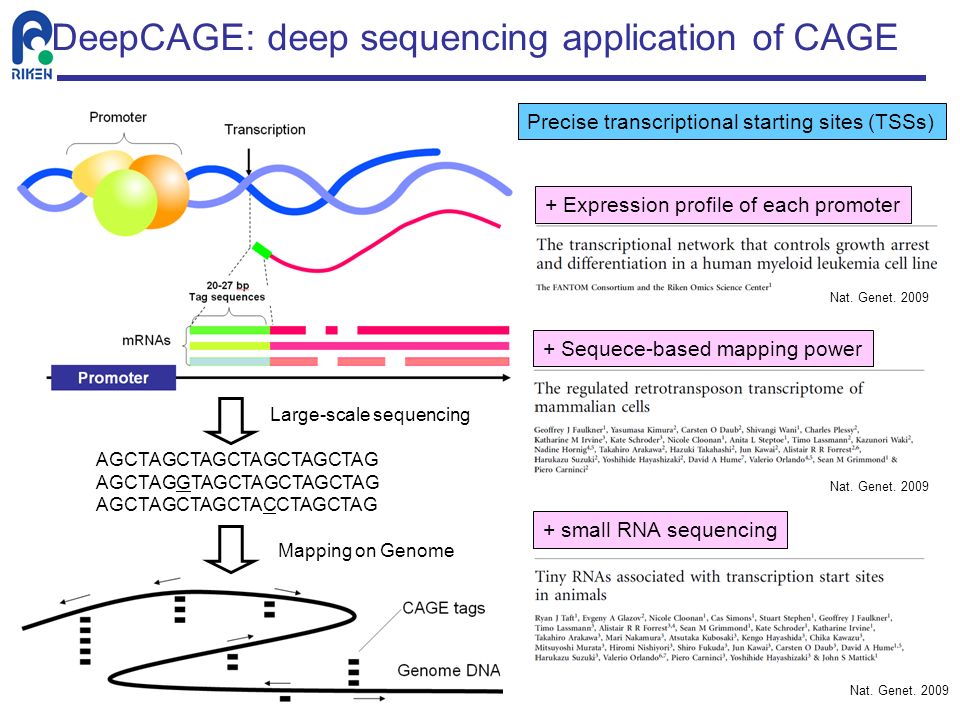
DeepCAGE: Incorporating Transcription Factors in Genome-wide Prediction of Chromatin Accessibility - ScienceDirect

DeepCAGE: Incorporating Transcription Factors in Genome-wide Prediction of Chromatin Accessibility - ScienceDirect
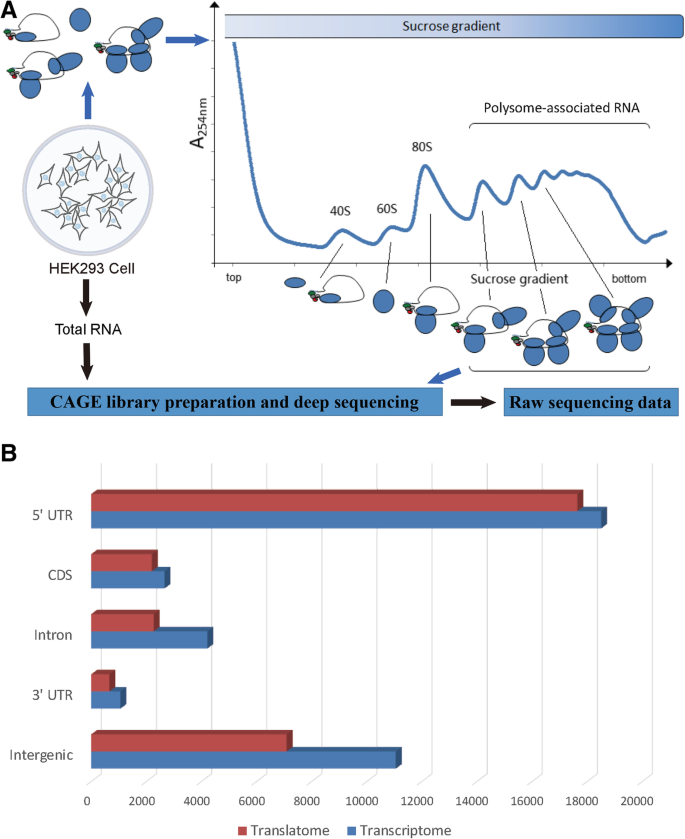
Selective translational usage of TSS and core promoters revealed by translatome sequencing | BMC Genomics | Full Text

Frontiers | Extensive Variation in Gene Expression is Revealed in 13 Fertility-Related Genes Using RNA-Seq, ISO-Seq, and CAGE-Seq From Brahman Cattle